AUREMOL Version 2.5.1
This time limited software version is free of charge (100 days after installation, no additional license required).
Permanent licenses can be obtained from Kalbitzer Innovations UG (kalbitzer-innovations@t-online.de).
The actual version (64 bit) is running under Windows 10.
When using the program please cite this site and/or
Gronwald, W. and Kalbitzer, H. R. (2004) Automated Structure Determination of Proteins by NMR Spectroscopy. Progr. NMR Spectr. 44, 33-96.
For citing specific methods see Programs.
PERMOL+
The program PERMOL+ for homology modelling of protein-protein and protein-ligand complexes. It is part of AUREMOL 2.5.1 but exists also as stand-alone version. It can be downloaded free of charge. Version running on Windows 10 or Mac OS are provided
When using the program please cite this site and Möglich, A., Weinfurtner, D., Maurer, T., Gronwald, W., Kalbitzer, H. R. (2005b) A restraint molecular dynamics and simulated annealing approach for protein homology modeling utilizing mean angles. BMC Bioinformatics, 6:91. Additional references see Programs.
UACSB
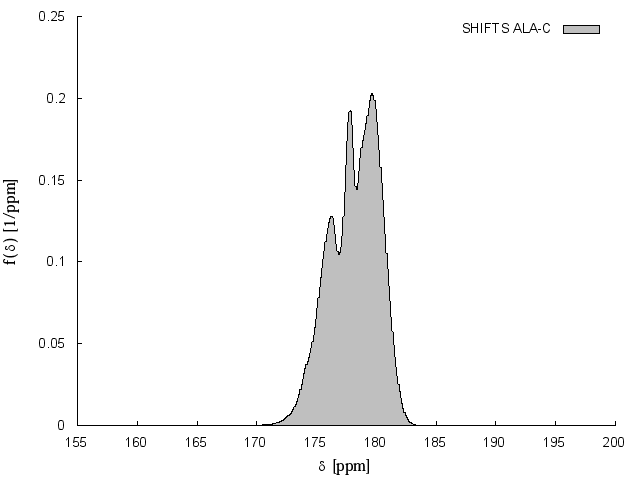
The unbiased artificial chemical shift data base (UACSB) was recalculated from the unbiased protein structural data base NH3D (Sillitoe et al., 2013). In the actual version, the chemical shift distributions of calculated from this database can be downloaded as pdf. A stand-alone version is actually not available.
IndirectRef
IndirectRef is a html-program for indirect referencing of heteronuclear NMR-spectra. It can be downloaded free of charge. When using the program please cite this site as well as the references given on the page Programs (IndirectRef).